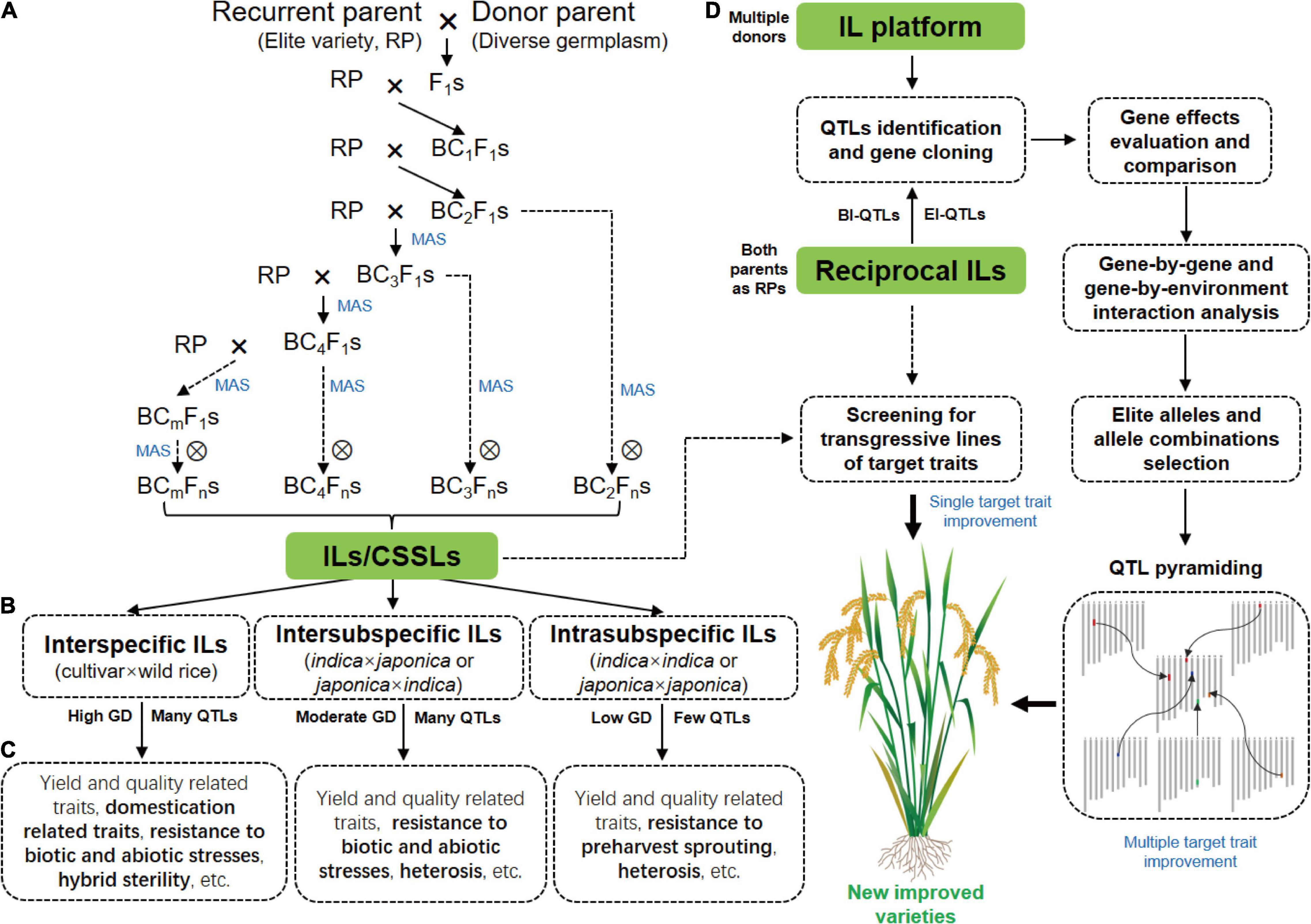
Software and Databases for managing and selecting molecular markers General introduction Pathway approach for candidate gene identification and introduction. - ppt download

Deep Learning Genome-wide Linkage Association Study for Wheat Fusarium Head Blight Resistance Genes Discovery | bioRxiv
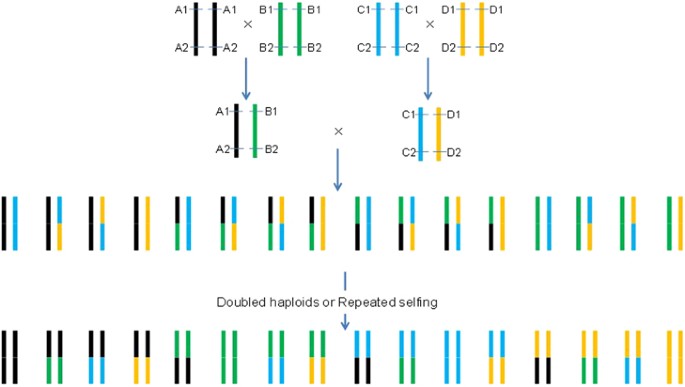
Background controlled QTL mapping in pure-line genetic populations derived from four-way crosses | Heredity

High-density SNP mapping reveals closely linked QTL for resistance to Stagonospora nodorum blotch (SNB) in flag leaf and glume of hexaploid wheat
DNA Marker Transmission and Linkage Analysis in Populations Derived from a Sugarcane (Saccharum spp.) x Erianthus arundinaceus Hybrid | PLOS ONE

PDF) Bridging the genotyping gap: using genotyping by sequencing (GBS) to add high-density SNP markers and new value to traditional bi-parental mapping and breeding populations | Susan McCouch - Academia.edu

Figure 1 | Biparental Crossing and QTL Mapping for Validation of Genome-Wide Association Studies | SpringerLink

Bridging the genotyping gap: using genotyping by sequencing (GBS) to add high-density SNP markers and new value to traditional bi-parental mapping and breeding populations
Genotyping-by-sequencing markers facilitate the identification of quantitative trait loci controlling resistance to Penicillium expansum in Malus sieversii | PLOS ONE
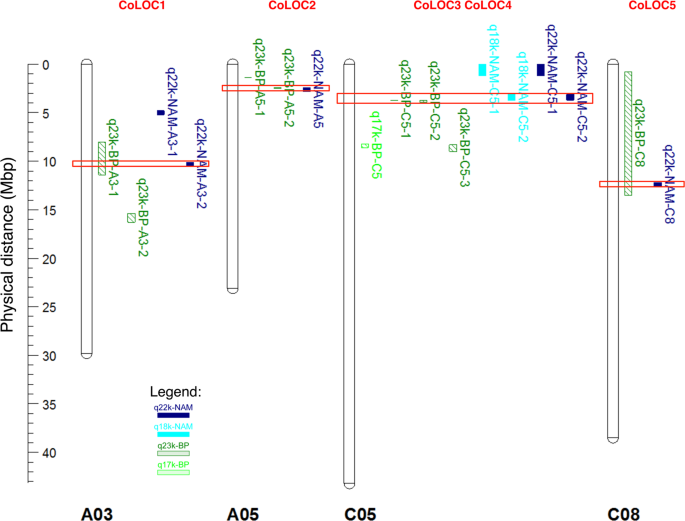
Gene presence-absence variation associates with quantitative Verticillium longisporum disease resistance in Brassica napus | Scientific Reports
Simple Sequence Repeat Marker Analysis of Genetic Diversity among Progeny of a Biparental Mapping Population of Sweetpotato
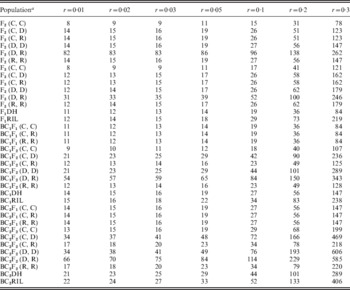
Estimation of recombination frequency in bi-parental genetic populations | Genetics Research | Cambridge Core
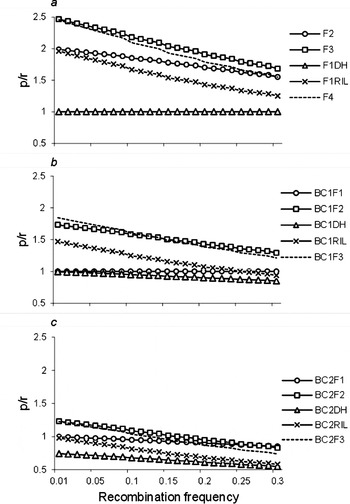
Estimation of recombination frequency in bi-parental genetic populations | Genetics Research | Cambridge Core
![PDF] Bi-parental cytoplasmic DNA inheritance in Wisteria (Fabaceae): evidence from a natural experiment. | Semantic Scholar PDF] Bi-parental cytoplasmic DNA inheritance in Wisteria (Fabaceae): evidence from a natural experiment. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/f85c9d5ad27183dd0a164e3e37b3b05c588a051c/2-Table1-1.png)
PDF] Bi-parental cytoplasmic DNA inheritance in Wisteria (Fabaceae): evidence from a natural experiment. | Semantic Scholar
Imputation of single nucleotide polymorphism genotypes in biparental, backcross, and topcross populations with a hidden Markov model
High-throughput sequencing techniques to flax genetics and breeding - Akhmetshina - Ecological genetics
A novel resistance gene for bacterial blight in rice, Xa43(t) identified by GWAS, confirmed by QTL mapping using a bi-parental population | PLOS ONE